Fluorescence Polarisation
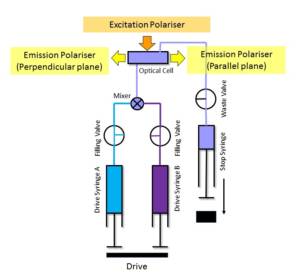
The fluorescence polarisation technique can be used with stopped-flow instruments to allow anisotropy measurements as well as total fluorescence measurements to be made. This is achieved by simultaneously detecting the fluorescence contributions of a sample from light polarised perpendicular and parallel to the parallel plane of the excitation beam. Using a fluorescence polarisation accessory, a near collimated excitation beam is delivered to the sample cell through a single fibre optic system.
This arrangement allows the use of a calcite prism polariser with transparency down to 220 nm. The sample cell is configured in a T-format for dual channel fluorescence detection and the photomultiplier couplings are fitted with UV dichroic sheet polarisers. The polarisers are arranged to pass light in the parallel and perpendicular planes with respect to the parallel plane of the excitation beam. The signals from each photomultiplier are processed and displayed using Kinetic Studio. The accessory (OPT-661) is available for use with all KinetAsyst dedicated stopped-flow fluorimeters.
In broad terms, fluorescence polarisation is relevant where a small (tumbling), fluorescent molecule binds a larger target molecule. Thus the observation of small fluorescent ligands binding protein molecules or short chain polymers (e.g. DNA) are often experiments suitable for this type of detection.
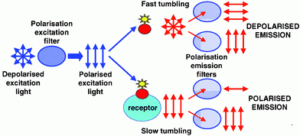
Fluorescence anisotropy is a method for measuring the binding interaction between two molecules, and can be used to measure the binding constant (or the inverse, the disassociation constant) for the interaction. The basic idea is that a fluorophore excited by polarized light (light whose “waves” only go one direction) will also emit polarized light. However, if a molecule is moving, it will tend to “scramble” the polarization of the light by radiating at a different direction from the incident light. The “scrambling” effect is greatest with fluorophores freely tumbling in solution and decreases with decreased rates of tumbling. Protein interactions can be detected when one of the interacting partners is fused to a fluorophore: upon binding of the partner molecule a larger, more stable complex is formed which will tumble more slowly (thus, increasing the polarization of the emitted light and reducing the “scrambling” effect). This technique works best if a small molecule is fused to a fluorophore and binds to a larger partner (this maximizes the difference in signal between bound and unbound states).
- AN.012.S60
Binding studies of a DNA repair enzyme using the SFA-20 Stopped-Flow fluorescence anisotropy
Keywords
- Fluorescence polarisation
- stopped-flow
- anisotropy
- DNA-protein interactions


